Annals of Neurosciences, Volume 19, Issue 2 (April), 2012
Development of time-resolved fluorescent based [EU]-GTP binding assay for selection of human Histamine 3 receptor antagonists/inverse agonist: a potential target for Alzheimer’s treatment
KEY WORDS
Alzheimer’s disease
Attention-deficit hyperactivity disorders
Histamine H3
Europium labeled Guanosine -5’
-triphosphate
HTS
ABSTRACT
Background: The histamine H3 receptor is an attractive G protein–coupled receptor drug target that regulates neurotransmission in the central nervous system and plays a crucial role in cognitive and homeostatic functions. This receptor exhibits molecular, pharmacological, and functional heterogeneity that affects the preclinical development of effective antagonists. The range of assay technologies like radio isotope based [35S] GTPγS binding assay, luminescent based reporter gene assay (In-direct cAMP measurement) for binding and signaling have been developed in High Throughput Screening (HTS) laboratories for the identification of hit or lead compounds acting on H3 receptor. Purpose: The [35S] GTPγS binding assay still remains a useful and a simple technique to demonstrate receptor activation and is one of the few functional, cell-free assays that has set the standards in the field of research. However, its radioactive nature imposes clear limitations to its use in regular laboratory practice and in high-throughput experimentation. Methods: Herein, we have developed and optimized a membrane based non-radioactive assay using a europium-labeled GTP analogue in which europium-GTP binding can be assayed using time-resolved fluorescence technology. Results: The characterization of H3 agonist or antagonist with HTRF platform has revealed a rank order potency (pEC50 & PKB) comparable to that from isotopic functional studies measured by liquid scintillation counter (LSC). Lastly, the Eu-GTP binding assay has been found to be highly robust (Z’ factor 0.84) with high percentage over basal counts. Conclusion: This assay can be utilized as a component of cascade for the screening of H3 receptor ligands.
doi : 10.5214/ans.0972.7531.12190205
*Corresponding Author:
Jitendra K. Singh
Tel : +91-2717656415
E-mail : jitendra@o2h.com
Introduction
The neurotransmitter Histamine exerts a wide range of biological functions, including neurotransmission, inflammation, smooth muscle contraction, dilation of capillaries and gastric acid secretion.1 These effects are mediated through four G-protein-coupled receptor subtypes2: h1, h2, h3 and h4. The h1 and h2 receptors are involved with the allergic response and gastric acid secretion, respectively.1 The h4 receptor is predominantly expressed in mast cells, and critically involved in the regulation of inflammatory and immune responses.3The Histamine h3 receptor was discovered in 1983 by Arrang and colleagues, using classic pharmacological approaches.4 It has been identified in both the central nervous system (CNS) as well as the peripheral nervous system as a presynaptic receptor controlling the release of histamine.4 It is also recognized as a heteroreceptor on non-histaminergic neurons that are capable of regulating the release of many other neurotransmitters, such as acetylcholine, nor-epinephrine, dopamine and serotonin.5
The h3 receptor is expressed predominantly in the central nervous system (CNS), with highest expression in the amygdala, substantia nigra, cerebral cortex and hypothalamus.6 These brain regions have been associated with cognition (cortex and hippocampus), sleep and homeostatic regulation (hypothalamus).7 The CNS effect of the H3 antagonists makes them valuable candidates for the treatment of obesity, epilepsy and age-related memory disorders, such as Alzheimer’s disease and attention-deficit hyperactivity disorders (ADHD).8 The histamine 3 receptor has been shown to couple with heterotrimeric Gi/o proteins and identification of several signal transduction pathways activated by the receptor:9 inhibition of adenylate cyclase, activation of phospholipase A2, activation of mitogen-activated protein kinase and Akt /glycogen synthase kinase-3β pathways, inhibition of calcium influx through voltage-gated calcium channel.
In this regard estimating GPCR activation by GTP binding is an important tool to study the early stage of signal transduction. Previously, this has been done using the radiolabeled non-hydrolyzable GTP analogue [35S] GTPγS.10 On the other hand, because of inherent limitations associated with working with radioactivity, there is an obvious advantage to adapting the GTP binding to a non- radioactive format.
Time-resolved fluorescence (TRF) technology exploits the unique fluorescence properties of Lanthanide chelates. It has a few advantages like low back ground and high-signal to noise ratios.11 In TRF, emission of the fluorescent label having a long decay time is recorded with a delay of about 0.4 ms after excitation, when the nonspecific fluorescence has died out. This results in an excellent signal-to-noise ratio offering extreme sensitivity, comparable to or even exceeding that of the radioactive compounds.12 The high-throughput format of the TRF assay makes the Eu-GTP-binding assay ideal for GPCR-targeted drug discovery13 and has been applied on membrane preparations containing various GPCRs such as α2A-Adrenergic,14 Neuropeptide,15 Dopamine,16 Muscarinic,17 and Chemokine receptors.18
In this present study, we describe the Eu-GTP binding assay for H3 receptor. We have employed Eu-GTP binding assay as part of our in vitro pharmacological characterization of novel h3 ligands. This assay protocol is also employed during each step in the development and optimization of a high-throughput assay. Typically, the assay development was started by determining the optimal receptor protein concentration followed by optimizing specific assay components (MgCl2 GDP, NaCl, Saponin and dimethyl sulfoxide (DMSO). Once the assay was developed, it was assessed for HTS compatibility by determining the critical factors reproducibility and signal to noise discrimination which govern the screening capability of the assay. Data generated using this assay is comparable with the [35S] GTPγS, with the advantages and convenience of a non-radioactive assay.
Methods
Chemicals
Histamine, R(-)-α-methylhistamine dihydrochloride, Imetit dihydrobromide, Immepip dihydro bromide, Clobenpropit dihydrobromide, Thioperamide maleate, GDP, Saponin, Ascorbic acid, Bovine serum albumin (BSA), Magnesium chloride and GTPγS were purchased from Sigma Aldrich (St. Louis, MO). Proxyfan oxalate was procured from Tocris Bioscience (UK). GSK 189254 was synthesized and provided by Oxygen Healthcare Research Private Limited. Eu-labeled GTP a fluorescent chelate was obtained from Perkin Elmer’s DELFIA Eu-GTP binding kit (product number AD0260) (Boston, MA). Human recombinant Histamine 3 receptor membrane protein was obtained from PerkinElmer. HPLC grade water was procured from Milli-Q water system (Millipore, USA). All other chemicals used in the study were of analytical grade and of proven purity.
Equipments
Analysis was performed on Victor3 ™ 1420 Multilabel counter (Perkin Elmer). Software for controlling the system was Wallac 1420 version 3.0. Filtration and analysis of the compounds was performed in Acrowell ™ 96 well filter membrane bottom plates (GHP membrane) were from Pall Life Sciences (USA). MultiScreenHTS Vacuum Manifold (Millipore) unit was used to filter out free Eu-GTP. The assay incubation was done in the Boekel Jitterbug microplate incubator shaker.
Histamine 3 Receptor Eu-GTP binding assay
The Eu-GTP binding assay was performed according to Frang et al.14 and as suggested by the manufacturer for the DELFIA Eu-GTP binding kit (Perkin Elmer). Human recombinant H3R membranes (1Unit, 15µg/mL.) were pre-incubated in the 96-well V bottom stock plate (Corning) for 15 minutes at 300C in final volume of 0.2 mL with 50 mM HEPES buffer, pH 7.4, supplemented with NaCl, MgCl2, Saponin, BSA, GDP and 50 µL of desired concentration of agonist in the wells. After pre-incubation for 15 minutes the reaction was started by addition of Eu-GTP to a final concentration of 3 nM and was further incubated for 30 minutes at 300C. After completion of the incubation the reaction mixture was transferred in Acrowell ™ 96 GHP filter plates (Pall). The assay was terminated by vacuum filtration (MultiScreen Vacuum Manifold, Millipore), followed by two washes with ice-cold buffer (50mM Tris-HCl, pH 7.4 & 10mM MgCl2). Plates were immediately allowed to Victor3™ 1420 Multilabel counter (Perkin Elmer Life Sciences) to measure the bound Eu-GTP with the membrane. The plate was read using the factory set protocol for europium measurements (340 excitation/ 615 nm emission, 0.4ms delay and 0.4 ms window). Nonspecific binding was determined by the inclusion of reagent listed above plus a final buffer concentration of 10 µM GTPγS.
Data Analysis
Data were measured as Europium GTP binding counts and percentage over basal level was calculated by using the following equation, the nonspecific binding (average) counts were subtracted from each data point:
% Over basal =
((Stimulated Signal (average)*100)/ (Basal Signal (average))-100
Inhibitor potency (EC50) was calculated using a 4-parameter, variable-slope, nonlinear iterative curve fitting provided by Prism® (GraphPad Software, Inc., San Diego, CA.) The EC50 values of individual agonists were calculated using the normalized percentage of maximum response.
The KB values of antagonists were calculated using the following equation:
KB = EC50/ (1+ [Ligand Concentration]/ Kd)
And the Z factor was calculated using the equation by Zhang et al [19].
Z’ factor =
1- ((3* SD of Max Signal + 3* SD of Min Signal)/ (Mean of Max Signal – Mean of Min Signal)
Where SD of Max Signal and SD of Min Signal are the corresponding standard deviations of Mean of Max Signal and Mean of Min Signal
Results and Discussion
The first stage of assay development was to determine the binding of Eu-GTP to membranes containing H3R to obtain significant difference between the basal binding activity and Imetit stimulated samples. Typically four independent experiments in triplicate were performed; the basal level with 3nM Eu-GTP was 6115 ± 231 (mean ± SD) counts in 30 minute, the stimulated level with 100 nM Imetit was 17776 ± 361.87 counts, and nonspecific binding was obtained in the presence of 10 µM GTPγS was 2238 ± 111 counts, resulting in 191± 5% over the basal level. This indicates the specific binding activity of the Eu-GTP to h3 Receptor coupled G proteins. Eu-GTP binding was also tested with control membrane protein (non transfected CHO cells). Membrane from non-transfected cells did not show any stimulation of Eu-GTP binding with 100nM Imetit, thus further confirming the specific binding of Eu-GTP.
Optimization of the Eu-GTP binding assay
The assay optimization steps were adapted from the PerkinElmer Delfia GTP binding protocol. The initial titration matrix for the each point was done in triplicate. A baseline control without agonist was included for each time point of the titration. A final concentration of 100 nM Imetit was used to activate the receptor.
Optimization of MgCl2 and GDP concentration
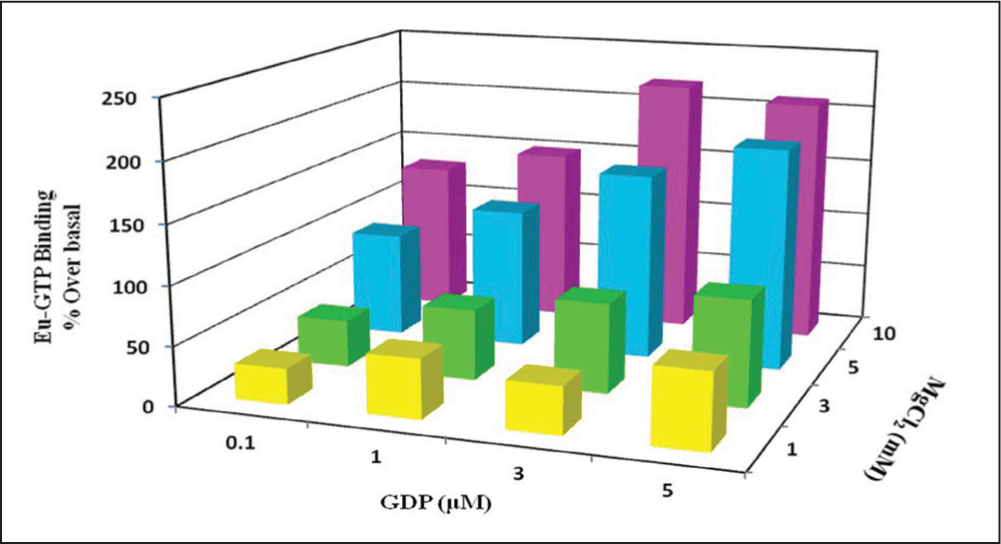
The concentration of MgCl2 and GDP was optimized by cross-titration on a 96-well Acro Well filter plate in concentration ranging from 0.1-10µM GDP and 1-10 mM MgCl2 as per manufacturer’s instructions. We have prepared concentration matrix with final assay concentration of 1, 3, 5 and 10 mM MgCl2 that was cross titrated with 0.1, 1, 3 and 5 µM GDP, keeping the NaCl concentration constant (100mM) in 50 mM HEPES buffer, pH 7.4 as reaction buffer for human h3 receptor. Figure 1 shows the comparison of the stimulation caused by 100 nM Imetit in different buffer conditions with membrane expressing h3 receptors. The maximum percentage over basal values was found with 3 µM GDP and GDP effect was increased with increasing MgCl2 concentration. Therefore, 3 µM GDP and 10 mM MgCl2 were selected for further assay development.
 |
 |
|
| Fig. 1: Optimization of the MgCl2 and GDP concentrations for the H3R Eu-GTP assay | Fig. 3: Optimization of Saponin concentration for the H3R Eu-GTP binding assay | |
 |
 |
|
| Fig. 2: Optimization of the NaCl concentration for H3R Eu-GTP binding Assay | Fig. 4: Effect of membrane protein concentration on % over basal values in the H3R Eu-GTP binding assay | |
Optimization of NaCl concentration
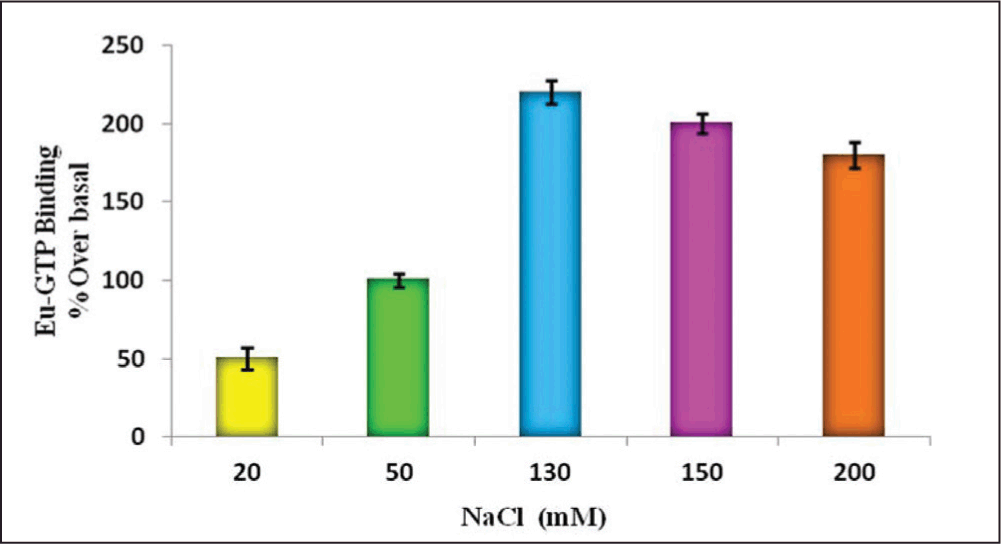
Similar protocol (used to optimize MgCl2 and GDP concentration) was used to optimize NaCl concentration in the assay. We selected 5 different concentrations of NaCl in the range from 20 to 200 mM. For this, NaCl stock solution was prepared with the recommended final assay concentration in 50 mM HEPES buffer, pH 7.4, by keeping the concentration of GDP (3µM) and MgCl2 (10mM) constant. The effect of Na+ concentration on stimulation of H3 receptor caused by 100nM Imetit is represented in Figure 2. The best percentage over basal ratio was found with 130 mM NaCl concentration.
Optimization of Saponin Concentration
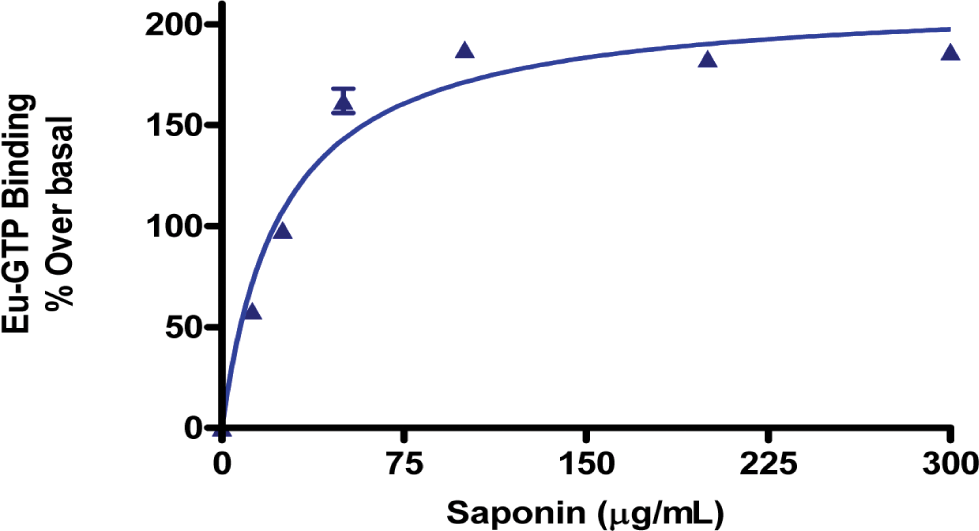
Saponin is a mild detergent, which can improve the GTP binding to h3 receptor, was tested within a range from 0-300 µg/mL. The stock solution of saponin with the recommended final assay concentration was made in 50 mM HEPES buffer pH 7.4 with the constant concentration of MgCl2 (10 mM), GDP (3 µM) and NaCl (130 mM). Figure 3 depicts the effect of saponin concentration on imetit induced binding of GTP-Eu to H3 receptor. The optimal signal was found with 100 µg/ml of saponin and this concentration was used in the subsequent steps of the assay development.
Optimization of h3 receptor membrane concentration in the binding assay
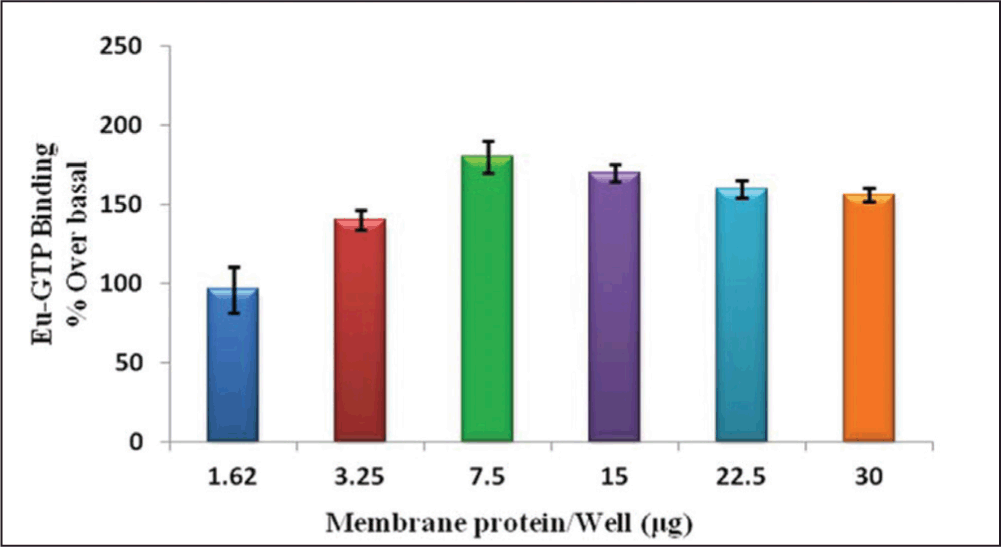
In all previous optimization steps, 15 µg (1Unit) of h3 receptor membrane per well was used, which is a fairly high amount of protein per well for the antagonist screening purpose. Therefore, we tested smaller protein amounts per well, and Figure 4 shows that even 1.62 µg provided reasonable percentage over basal (96%) values compared with 15 µg per well (170). We have performed the binding assay to obtain optimal % over basal, upto 30 µg per well and the best % over basal ratio was found with 7.5 µg/well.
Validation of the Eu-GTP binding assay using known Agonists
To further validate the Eu-GTP binding assay, five histamine 3 receptor agonist previously screened in a radiometric ([35S] GTPγS) assay were tested. These agonists were selected with a wide range of affinity, which includes pEC50 value from low nM to High nM. As shown in the Table 1, Figure 5 the pEC50 values for the 5 agonists were very similar to those determined in the isotopic [35S] GTPγS functional assay.19
DMSO tolerance study
To adapt the assay for antagonist selection, a DMSO tolerance study was performed. The H3R activity was not significantly affected by up to 3% DMSO (data not shown). The antagonist screening was then conducted at 2% DMSO.
Table 1: Dose-response isotherms of ligands were analyzed by non-linear regression. Agonist potency is expressed as pEC50. The antagonist potency to inhibit Eu-GTP binding induced by 100nM Imetit is expressed as PKB. Results are presented as means ± S.E.M. of at least three independent experiments
| Ligand | Eu-GTP | [35S]GTPγS [20, 21] |
| Agonists |
| Imetit | 9.03 ± 0.11 | 8.82 ± 0.07 |
| Immepip | 9.44 ± 0.01 | 9.29 ± 0.07 |
| R(-)-α-methylh- istamine | 8.85 ± 0.08 | 8.57 ± 0.01 |
| Proxyfan | 8.36 ± 0.12 | 8.3 ± 0.10 |
| Histamine | 7.55 ± 0.07 | 7.46 ± 0.04 |
| Antagonists |
| GSK189254 | 9.90 ± 0.11 | 9.06 ± 0.02 |
| Clobenpropit | 9.00 ± 0.02 | 9.28 ± 0.06 |
| Thioperamide | 8.26 ± 0.10 | 8.12 ± 0.01 |
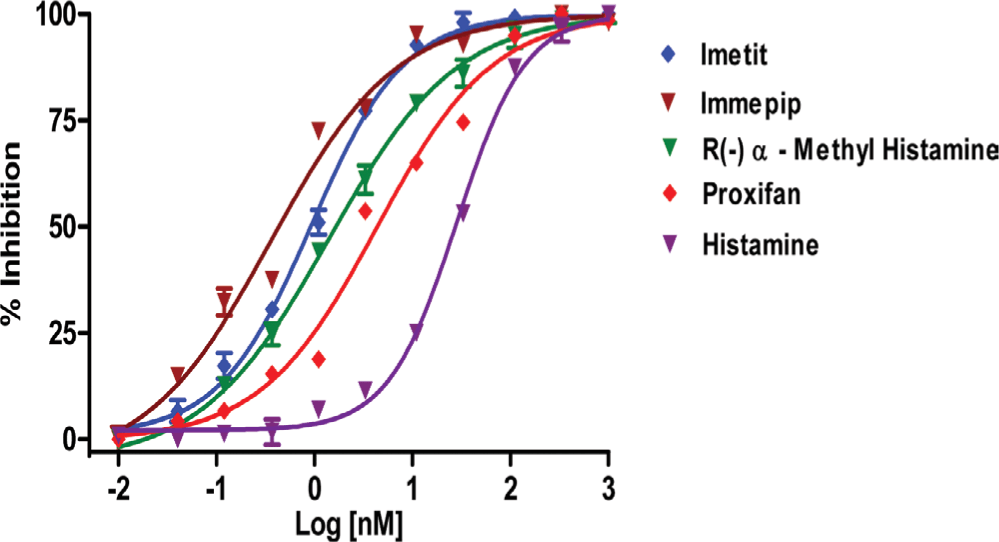
Fig. 5: Dose –response curve of agonist-induced binding of Eu-GTP to H3R, investigated with 5 different H3R agonists
Adaptation of Eu-GTP binding assay for h3 receptor Antagonists selection
The aim of these modifications to the assay was to use it to screen the h3 receptor antagonists. In order to utilize the assay for this purpose the h3 receptor membranes were pre-incubated with agonists and antagonists for 15 minutes before the addition of Eu-GTP in the 96-well V bottom assay plate, by keeping other components constant. The potencies of h3 receptor antagonists (GSK 189254, Clobenpropit and Thioperamide) were determined for antagonism of Imetit- mediated inhibition of Eu-GTP binding to human h3 receptor. The data was analyzed in nonlinear regression mode to obtain PKB values by using Graph pad Prism software which is shown in Table 1, Figure 6. These values are consistent with previously published data.20
Z’ Determination
To be of use in agonist and antagonist screening, the assay must fulfill other criteria in that it must be reproducible, rendering data comparable to the gold standard radiometric assay, and be free of interference from test compounds. To address these points, the Z’ value for the assay, the standard measure of reproducibility and signal-to-noise discrimination, were determined.19 The Z’ value is obtained by conducting background and maximum stimulation for multiple times across assay plate on separate days. The data shown in Figure 7 gives calculated Z’ factor of 0.847. These excellent scores indicate that this assay is viable for use in high throughput screening.
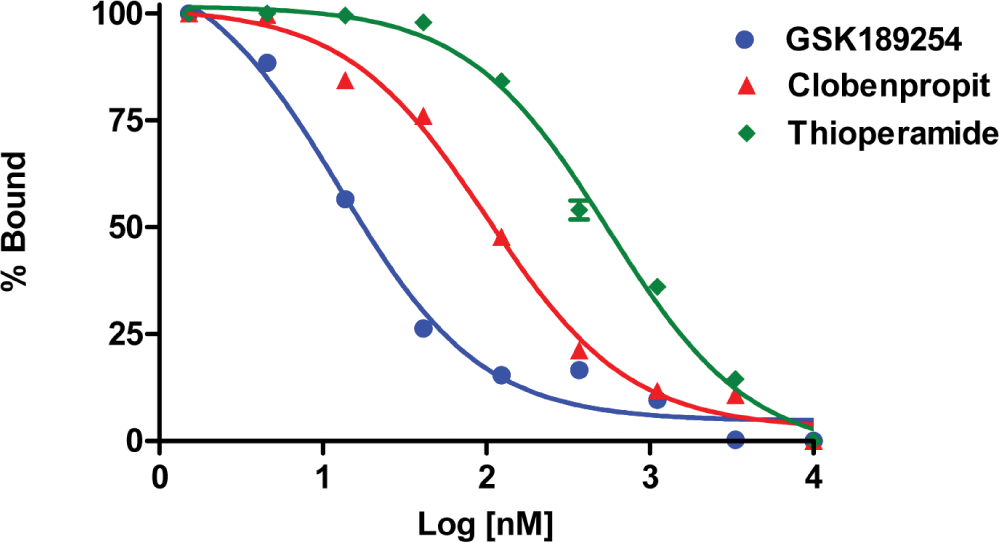
Fig. 6: Antagonism of Imetit-stimulated Eu-GTP binding: [Antagonist concentration-inhibition curves for antagonism of 100 nM Imetit responses by GSK 189254, Clobenpropit and Thioperamide.] The data shown are the average of at least three experiments (± SEM), each with three determinations per condition

Fig. 7: Z’ factor estimation for Eu-GTP binding assay
Conclusion
The data presented shows that time resolved fluorescence (TRF) based Eu-GTP binding assay is an excellent format for the selection of H3R agonist and antagonists in that it provides a simple homogeneous format that is more affordable, less time consuming, amendable to automation and scalable to any assay volume required. However, identification of appropriate assay for the characterization of agonist and antagonist of histamine 3 receptor presents a challenge for the GPCR functional assay development and impedes progression of the target in drug discovery program. Although, radioactive assay ([35S] GTPγS) had been developed for the selection of H3R agonist and antagonist, disposal costs and potential for the contamination of HTS facilities restricts the use of radioactive binding assay for screening purpose. The assay has an advantage over the radiometric assay, as it has long fluorescence half life and can detect less than one attomole of europium in a multiwall plate sample.
The assay, when compared with the gold standard radiometric formats for histamine 3 receptor functional assay, gives comparable Z’ values and in actual screening trials produces nearly identical results to those of radiometric assay.
Article complies with International Committee of Medical Journal editor uniform requirements for the manuscripts.
Competing interests: None, Source of Funding: House funding
Received Date : 21 July 2011; Revised Date: 2 February 2012
Accepted Date : 7 March 2012
Reference
1. Hill SJ, Ganelli CR, Timmerman H, et al. International Union of Pharmacology. XIII. Classification of histamine receptors. Pharmacol. Rev. 1997; 49: 253–278.
2. Parsons ME and Ganellin CR. Histamine and its receptors. Br. J. Pharmacol. 2006; 147: S127–S135.
3. De Esch IJP, Thurmond RL, Jongejan A, et al. The histamine H4 receptor as a new therapeutic target for inflammation. Trends Pharmacol. Sci 2005; 26: 462–469.
4. Arrang JM, Garbarg M and Schwartz J C. Auto inhibition of brain histamine release mediated by a novel class (H3) of histamine receptor. Nature 1983; 302: 832–837.
5. Schlicker E, Malinowska B, Kathmann M, et al. Modulation of Neurotransmitter Release via Histamine H3 Heteroreceptors. Fundam. Clin. Pharmacol.1994; 8: 128–137.
6. Drutel G, Peitsaro N, Karlstedt K, et al. Identification of rat H-3 receptor isoforms with different brain expression and signaling properties. Mol. Pharmacol. 2001; 59: 1–8.
7. Martinezmir MI, Pollard H, Moreau J, et al. Histamine receptor-H1, receptor-H2 and receptor-H3 visualized in the brain of human and nonhuman-primates. Brain Res.1990; 526: 322–327.
8. Leurs R, Blandina P, Tedford C, et al. Therapeutic potential of histamine H3 receptor agonists and antagonists. Trends Pharmacol. Sci.1998; 19: 177–183.
9. Bongers G, Bakker RA and Leurs R. Molecular aspects of the histamine H3 receptor. Biochem. Pharmacol. 2007; 73: 1195–1204.
10. Milligan G. Principles: extending the utility of [35S] GTPgammaS binding assays. Trends Pharmacol. Sci. 2003; 24: 87–90.
11. Selvin PR. Principles and biophysical applications of lanthanide-based probes. Annu. Rev. Biophys. Biomol. Struct. 2002; 31: 275–302.
12. Koval A, Kopein D, Purvanov V, et al. Europium-labelled GTP as a general nonradioactive substitute for [35S] GTPγS in high-throughput G protein studies. Anal. Biochem. 2010; 397: 202–207.
13. Hemmila IA and Hurskainen P. Novel detection strategies for drug discovery. Drug Discov. Today 2002; 7: S150–S156.
14. Frang H, Mukkala VM, Syysto R, et al. Nonradioactive GTP binding assay to monitor activation of g protein-coupled receptors. Assay Drug Dev. Technol. 2003; 1: 275–280.
15. Engstrom M, Narvanen A, Savola, JM, et al. Assessing activation of the human neuropeptide FF2 receptor with a non-radioactive GTP binding assay. Peptides 2004; 25: 2099–2104.
16. Leopoldo M, Lacivita E, Colabufo NA, et al. First structure-activity relationship study on dopamine D3 receptor agents with N-[4-(4-arylpiperazin-1-yl)butyl] arylcarboxamide structure. J. Med. Chem. 2005; 48: 7919–7922.
17. Zhang HY, Watson ML, Gallagher M, et al. Muscarinic receptor mediated GTP-Eu binding in the hippocampus and prefrontal cortex is correlated with spatial memory impairment in aged rats. Neurobiol. Aging 2007; 28: 619–626.
18. Labrecque J, Wong RSY and Fricker SPA. Time-resolved fluorescent lanthanide (Eu)-GTP binding assay for chemokine receptors as targets in drug discovery, Methods Mol. Biol. 2009; 552: 153–169.
19. Zhang JH, Chung TDY and Oldenburg KR. A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J. Biomol. Screen. 2000; 4: 67–73.
20. Coge F, Guenin SP, Audinot V, et al. Genomic organization and characterization of splice variants of the human histamine H3 receptor. Biochem. J.2001; 355: 279–288.